11 Oct 2022
Optimising pore size and particle type for biomolecule analysis
By Claire Pashley
Whilst stationary phase selection is a critical column parameter for small molecules, the impact of differing stationary phases for larger biomolecules is less so. [1] This is because the chromatographic selectivity of biomolecules is driven more by its solubility in the mobile phase rather than its propensity to interact with the stationary phase. [1] Although selectivity differences can be achieved using different stationary phase chemistries, usually this is controlled and optimised by adjusting the mobile phase gradient. [1] Therefore, when choosing a column for biomolecule analysis, other column parameters such as pore size and particle type become far more important to consider for optimal chromatography.
Demystifying reversed phase column
selection for biomolecule analysis
Take the quick guide on CHROMacademy
Pore size selection
The overall size of an analyte in solution will determine whether it can obtain unrestricted access into and out of the pores in a chromatography column [2]. Since most of the bonded phase and surface area reside in the pores, a pore size that is too small will reduce retention of the analyte on the column. Furthermore, restricted mass transfer (diffusion in and out of the pores) contributes to peak dispersion and can result in broader peaks [2]. Since biomolecules can have different shapes and sizes in solution even for the same molecular weight (MW) careful consideration needs to be given to a molecules hydrodynamic radius when selecting a column for HPLC analysis. [2] However, a consensus has not yet been reached on how large pore size should be (relative to molecular size) to avoid unnecessary loss of chromatography performance. Some have suggested that the particle pore size should be a minimum of 8-10 fold larger than the molecular diameter for minimal size exclusion effects and better access to the stationary phase and less restricted diffusion for large analytes. However, when interpreting pore size, it is worth remembering that the particles for UHPLC and HPLC usually have a broad pore size distribution range (shown in Figure 1), therefore a particular pore size may be relevant for a range of molecular sizes. For example, in Figure 1 there are recommendations for the minimum Halo® pore size for different molecules based on the pore distributions shown. Each manufacturer will have different pore size distributions for their silica and be able to make similar recommendations accordingly.
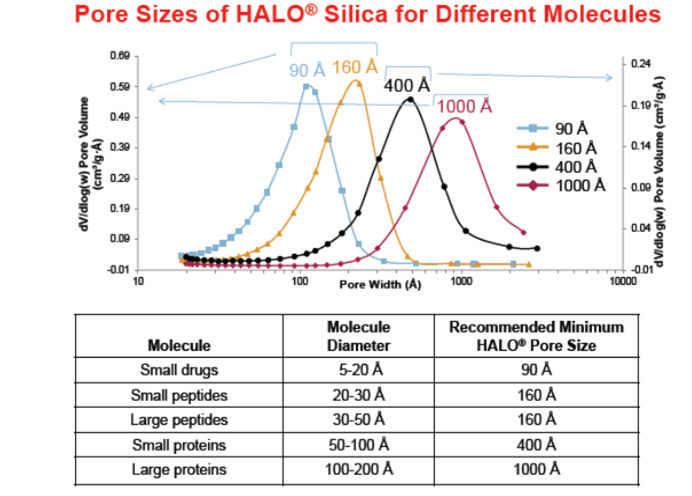
Figure 1: Halo pore size selection based on analyte size and type. (Figure taken from Reference [2])
When comparing columns from different manufacturers you will notice that 300 Å is often the “go-to” size for peptides and proteins. This has historically been the case and so many chromatographers automatically assume that this pore size will be suitable for large molecules, and they will not have any problems due to restricted pore access, slow diffusion, and size exclusion effects. [2] In many cases this is correct and such columns can give excellent resolution for large molecules such as monoclonal antibodies (mAbs), especially when an appropriately hydrophobic stationary phase chemistry and high temperature are used, as shown in Figure 2. However, for molecules greater than 50 kDa in size high resolution separation may be obtained using a larger pore size column (greater than 400 Å).

Method details
Column: YMC-Triart Bio C4 (3µm, 300 Å) 150 x 3.0 mm ID
Eluent: A) water/TFA (100/0.1)
B) acetonitrile/TFA (100/0.1)
Gradient: 0 min – 30 % B
15 mins – 60 % B
30 mins – 90 % B
Flow rate: 0.4 mL/min
Detection: UV at 220 nm
Injection: 4 µL (0.5 mg/mL)
Figure 2: Reverse Phase analysis of Bevacizumab on a 300 Å pore size column at different temperatures (Figure taken from Reference 3)
Particle size and morphology
In addition to optimising pore size, considering particle size and morphology is important to achieving good chromatography resolution. Large molecules have poor mass transfer kinetics and move in and out of the pores in the stationary phase slowly which results in broad peaks. [1] There are two ways to minimise the effects of slow mass transfer:
1) Use a traditional fully porous (FPP) stationary phase with a small particle size (< 2µm) which will have shallower pores than a larger particle
2) Use a superficially porous (SPP)
Using a superficially porous particles rather than standard fully porous particles can be worth considering when analysing large biomolecules. [4] These particles have a solid silica core and are surrounded by a thin, porous shell with differing pore sizes available (Figure 3). Using a combination of a wide pore size (i.e ≥ 400 Å) and thin shell mean that biomolecules can diffuse very quickly in and out of the pores and do not suffer from poor mass transfer, which may result in a more efficient application that provides sharper chromatography peaks. [4]
There are several advantages of using SPPs over small FPPs for biomolecule separations. Lower backpressures are generated so longer and/or narrower columns can be used. [1] Larger particle sizes can be used which are compatible with standard HPLC rather than UHPLC [1]. When using larger particle sizes, the columns will have larger interstitial gaps between the particles and are less prone to blockages compared to smaller particle columns. [1] SPPs offer improved peak efficiency compared with the equivalently sized FPP, therefore often offering better chromatographic performance. [1]
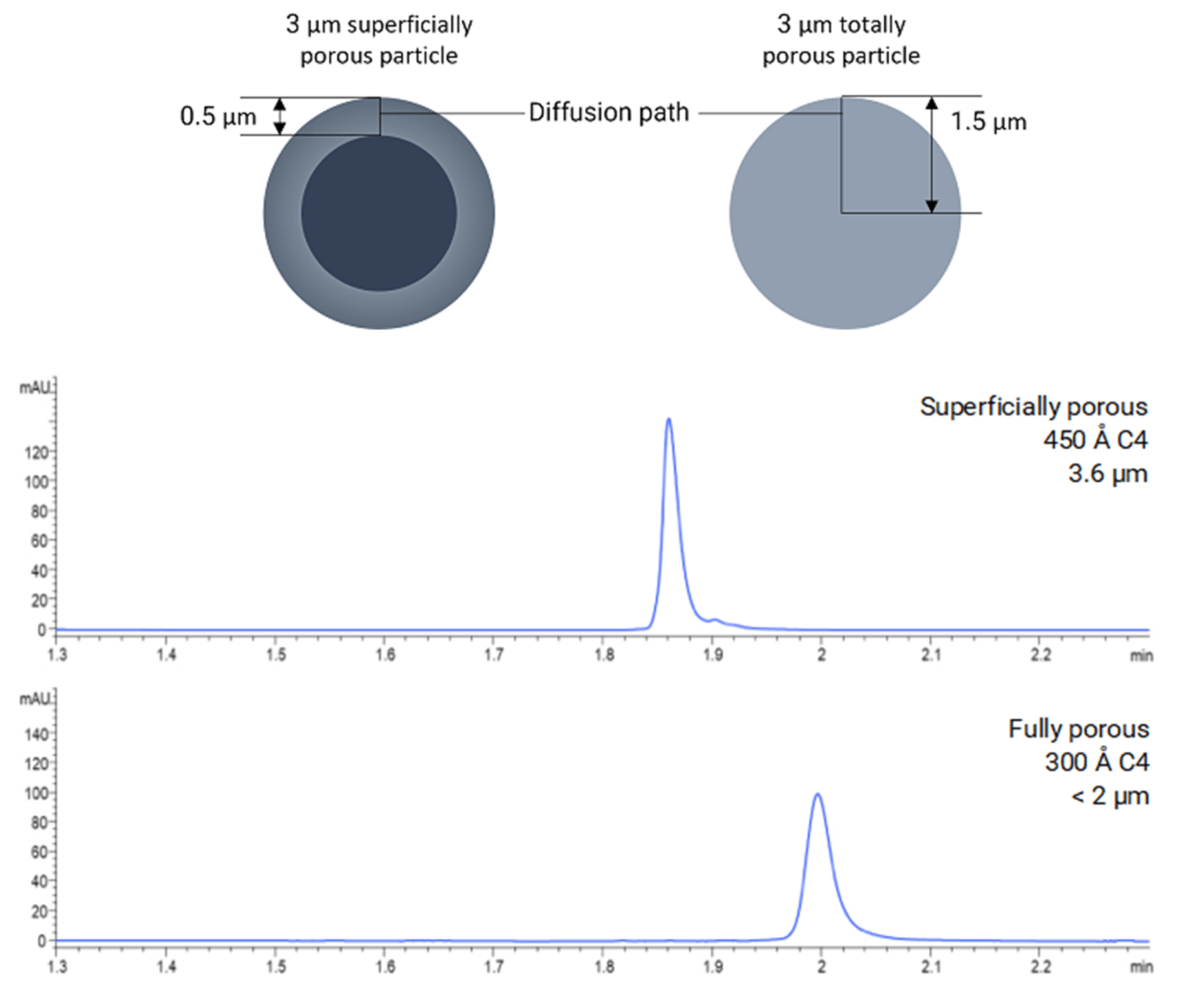
Figure 3: Effect of particle morphology on retention, efficiency, and resolution in an intact mAb analysis. Figure taken from Reference [1]
Summary
Given the column parameters discussed above, where possible it is sensible to evaluate both different pore size and particle type on your method, particularly if you are not getting the required resolution on a 300 Å FPP column. Figure 4 demonstrates the changes in separation of a mAb, with a molecular weight of about 150 kDa. In this case a larger pore size of 1000 Å, combined with a superficially porous particle, offers improved retention, sharper peaks and improved resolution of minor components compared to the 300 Å fully porous particle.
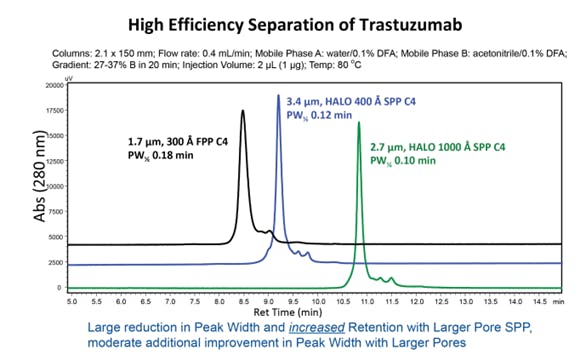
Figure 4: Comparison of 300, 400 and 1000 Å columns for reverse phase analysis of 148 kDa mAb (Figure taken from Reference 2)
References
[1] CHROMacademy lesson: Demystifying Reversed Phase Column Selection for Biomolecule Analysis: https://www.chromacademy.com/channels/biochromatography-training-courses/technique/demystifying-reversed-phase-column-selection-for-biomolecule-analysis/
[2] https://halocolumns.com/wp-content/uploads/2021/04/AMT-UTH-Pore-Size_BioClass_04_020.pdf