14 May 2019
Spectral Deconvolution
I was planning to write something about that momentous achievement of food science — the Chocolate Digestive… but somebody showed me a chromatogram last week that completely changed my mind. There was a distorted peak that looked like a coeluting impurity, but it was difficult to tell. I immediately said “that’s a job for AMDIS!” to which the other person replied “who is Andy?”
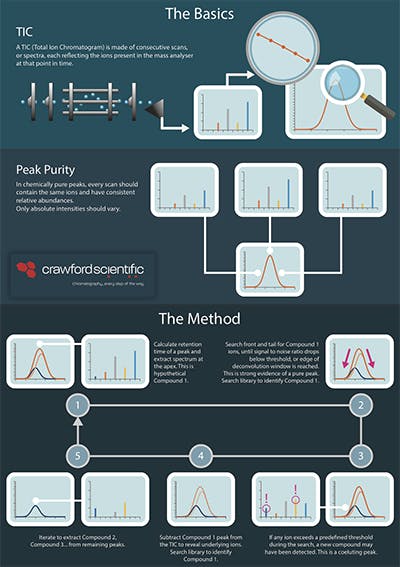
I quickly realised that many people may have never used deconvolution software, or really know how it works, and that’s how this infographic came to be. Because they should. But before we get to that, a little more background.
Peak shoulders, split peaks, and rider peaks are all common problems in separation sciences. With careful inspection of the offending peak shapes, we can guess that we are in front of two or more coeluting compounds, but it is impossible to say with confidence where one begins and the other ends.
Scanning detectors, like DAD or MS, offered a solution to the problem. With the spectra produced by these devices and clever data manipulation, one can unscramble the overlapping peaks. This procedure is known as spectral deconvolution.
Pioneering research in the early seventies led the way to the Automated Mass Spectral Deconvolution and Identification System, AMDIS (again, not Andy’s). Generations of GC-MS practitioners have used AMDIS, and it has been the template for subsequent deconvolution tools. We wanted to pay homage to this landmark programme, with an infographic that explains what AMDIS does and how it works.
Click here - or on the image below - for a full sized pdf